Projects
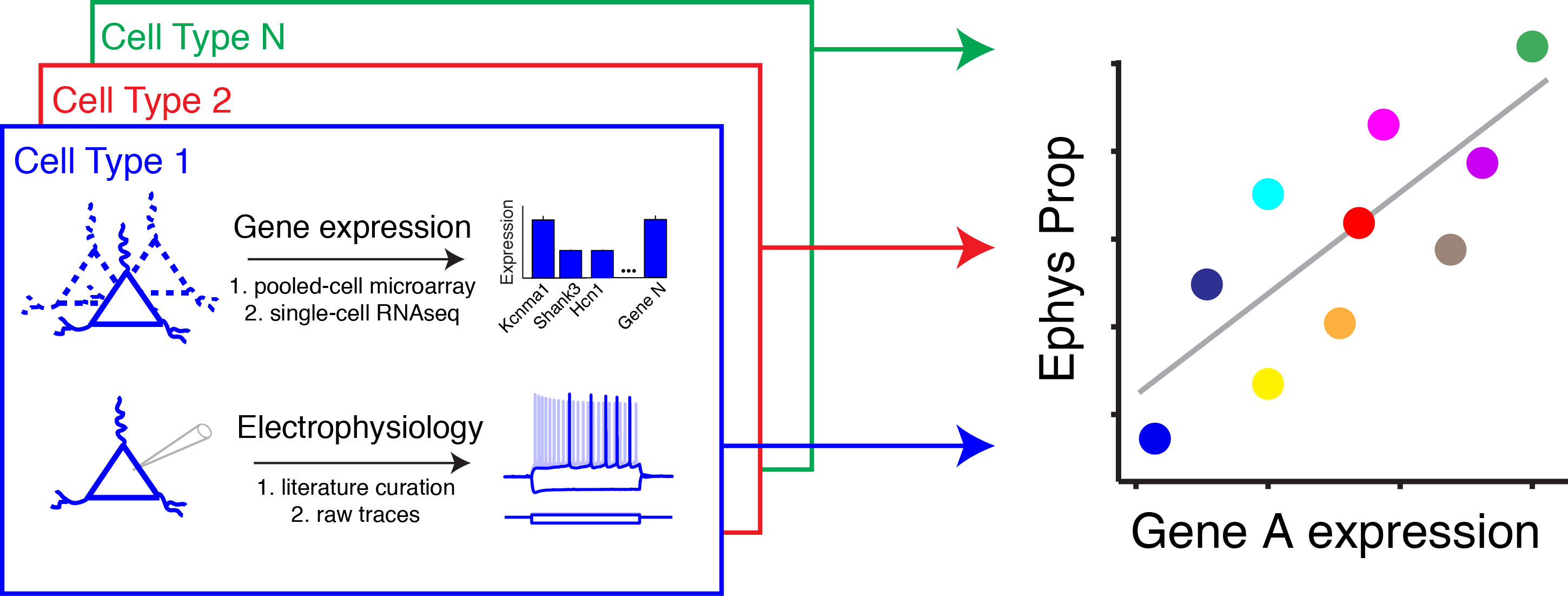
Linking gene expression to neuron function
We are developing machine learning methods to map the relationship between cell type-specific gene expression and phenotypic features of neuron function, including neuron electrophysiology and morphology.
Relevant publications: Körner, Howard et al., Cell 2026, Bomkamp & Tripathy et al., PLoS Comp Bio 2019
Relevant publications: Körner, Howard et al., Cell 2026, Bomkamp & Tripathy et al., PLoS Comp Bio 2019
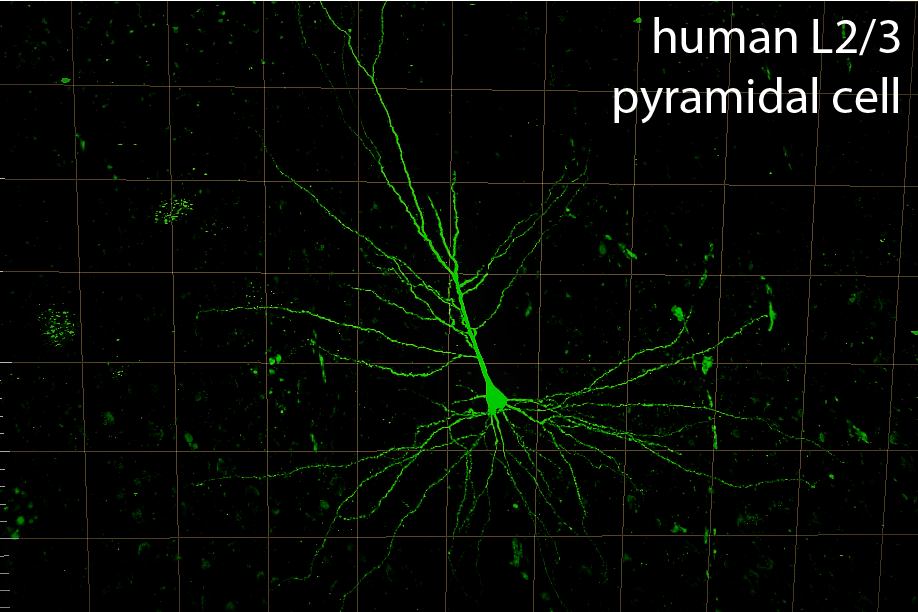
Studying the diversity and function of human and primate cortical neurons
We investigate the electrophysiological function of human and primate cortical neurons. In humans, we study tissue resected during routine neurosurgery in collaboration with the Valiante Lab at the Krembil Research Institute, correlating variability in neuron function to an individual’s genetics and demographic features. In non-human primates, we collaborate with the Martinez-Trujillo Lab to understand how single neuron diversity supports functional specialization across cortical areas.
Relevant publications: Moradi Chameh et al., Nat Comms 2021, Koch et al., Cell Reports 2025, and Feyerabend et al., bioRxiv 2024
Relevant publications: Moradi Chameh et al., Nat Comms 2021, Koch et al., Cell Reports 2025, and Feyerabend et al., bioRxiv 2024
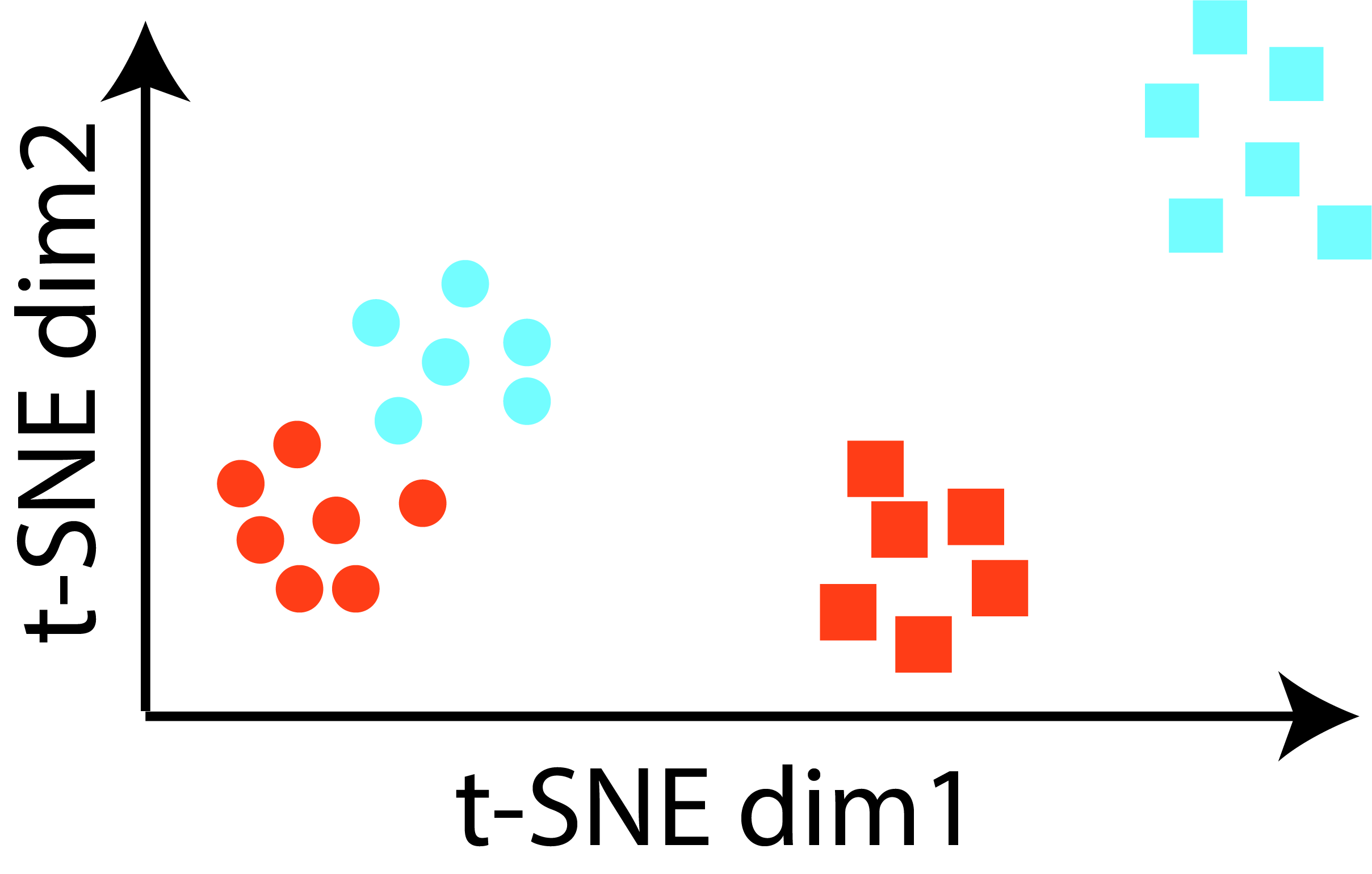

Using single-cell and spatial transcriptomics to understand brain disorders
Single-cell RNA-sequencing is revolutionizing our understanding of the cell types of the nervous system.
In collaboration with a number of experimental colleagues, we are developing computational methods to use these technologies to understand how
brain cells change in their proportion, gene expression, and electrophysiology due to neuropsychiatric and neurological conditions,
including depression, aging, and Alzheimer's disease.
Relevant publications: Arbabi et al., Mol Psychiatry 2024 and Kiss et al., Biol Psych: GOS 2026
Relevant publications: Arbabi et al., Mol Psychiatry 2024 and Kiss et al., Biol Psych: GOS 2026
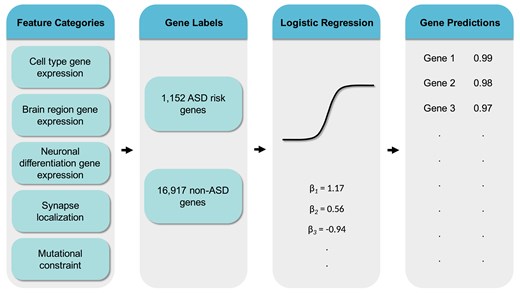
Genomics of neurodevelopmental disorders
We study the genetic architecture of neurodevelopmental conditions like autism spectrum disorder. This work includes identifying risk genes from consanguineous populations, building machine learning models to predict new ASD risk genes, and using long-read sequencing to characterize transcript isoform diversity during human neuronal differentiation.
Relevant publications: Harripaul et al., Sci Reports 2026, Rynard et al., Genetics 2025, and Xu et al., bioRxiv 2025
Relevant publications: Harripaul et al., Sci Reports 2026, Rynard et al., Genetics 2025, and Xu et al., bioRxiv 2025

Spatial transcriptomics of brain development and disease
We use spatial transcriptomic technologies to understand how brain cell types are organized in tissue and how their spatial relationships change in development, injury, and disease. This work spans perinatal brain injury, maternal brain plasticity, and the astrocyte response to stroke.
Relevant publications: Tahmasian et al., eNeuro 2024, Arbabi et al., bioRxiv 2026, and Scott et al., Nat Comms 2024
Relevant publications: Tahmasian et al., eNeuro 2024, Arbabi et al., bioRxiv 2026, and Scott et al., Nat Comms 2024
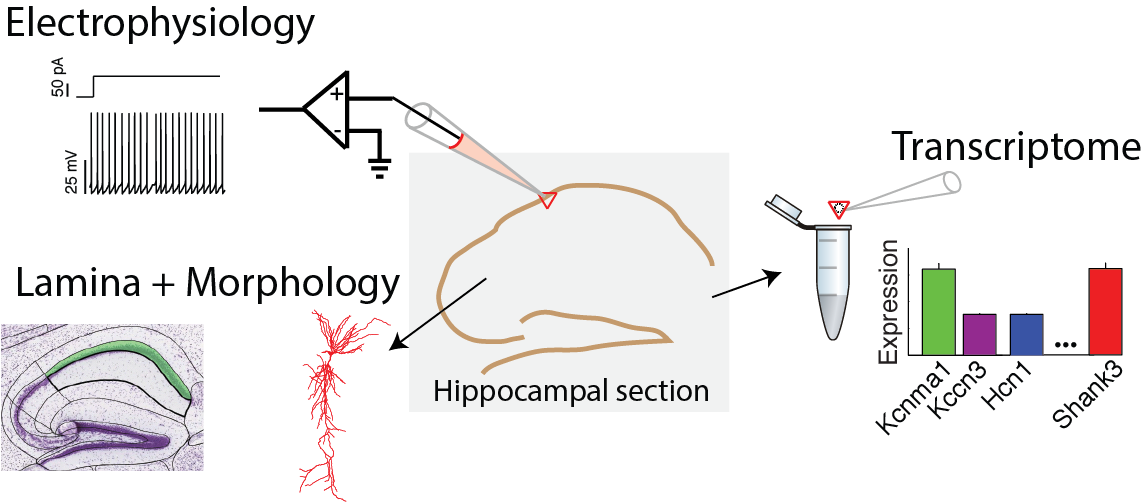
Neuroinformatics and methods for brain cell type characterization
We build public databases on brain cell type diversity, including
NeuroElectro.org and
NeuroExpresso.org (in collaboration with the Pavlidis Lab at UBC),
and develop computational methods to assess and improve the quality of multimodal single-cell datasets like Patch-seq.
Relevant publications: Tripathy et al., J Neurophysiol 2015, Mancarci et al., eNeuro 2017, Tripathy et al., Front Cell Neurosci 2018, and Arbabi et al., iScience 2023; YouTube video for NeuroElectro: here
Relevant publications: Tripathy et al., J Neurophysiol 2015, Mancarci et al., eNeuro 2017, Tripathy et al., Front Cell Neurosci 2018, and Arbabi et al., iScience 2023; YouTube video for NeuroElectro: here
Using large-scale population cohorts to understand the interplay between genetics, the brain, and psychiatric illnesses
We have begun using datasets like the UK Biobank
to better understand the multi-modal and multi-faceted aspects of psychiatric illness. We are using these datasets to pursue questions like:
"How is depression defined into subtypes?" and "How do genetics and environmental
influences, like poor sleep quality or cannabis use, interact to increase the risk of psychiatric disorders?".
Relevant publications: Wainberg et al., Nat Comms 2024 and Wainberg et al., JAMA Psychiatry 2022
Relevant publications: Wainberg et al., Nat Comms 2024 and Wainberg et al., JAMA Psychiatry 2022